-Search query
-Search result
Showing 1 - 50 of 602 items for (author: han & sj)

EMDB-19477: 
Saccharomyces cerevisiae FAS type I
Method: single particle / : Mann D, Grininger M, Ludig D, Sachse C

EMDB-19489: 
Tobacco mosaic virus from scanning transmission electron microscopy at CSA=2.0 mrad
Method: helical / : Mann D, Filopoulou A, Sachse C

EMDB-41907: 
Computationally Designed, Expandable O4 Octahedral Handshake Nanocage
Method: single particle / : Weidle C, Borst A
EMDB-42031: 
Computational Designed Nanocage O43_129_+8
Method: single particle / : Weidle C, Kibler RD

EMDB-43658: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43659: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43660: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-17171: 
CTE typeI tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17173: 
CTE typeIII tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17174: 
CTE typeII tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17175: 
TMEM106B Fold1-s filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17176: 
TMEM106B Fold I-d filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17177: 
Ab typeII filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17178: 
CTE typeI tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17179: 
TypeII tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-17180: 
CTE typeIII tau filament
Method: helical / : Tetter S, Qi C, Ryskeldi-Falcon B, Scheres SHW, Goedert M

EMDB-17181: 
PHF tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
PDB-8ot6: 
CTE typeI tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

PDB-8ot9: 
CTE typeIII tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
PDB-8otc: 
CTE typeII tau filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
PDB-8otd: 
TMEM106B Fold1-s filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

PDB-8ote: 
TMEM106B Fold I-d filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

PDB-8otf: 
Ab typeII filament from Guam ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
PDB-8otg: 
CTE typeI tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
PDB-8oth: 
TypeII tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M
PDB-8oti: 
CTE typeIII tau filament
Method: helical / : Tetter S, Qi C, Ryskeldi-Falcon B, Scheres SHW, Goedert M
PDB-8otj: 
PHF tau filament from Kii ALS/PDC
Method: helical / : Qi C, Yang S, Scheres SHW, Goedert M

EMDB-43318: 
Twistless helix 12 repeat ring design R12B
Method: single particle / : Calise SJ, Kollman JM
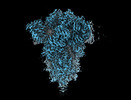
EMDB-18807: 
SD1-2 fab in complex with SARS-COV-2 BA.12.1 Spike Glycoprotein.
Method: single particle / : Duyvesteyn HME, Ren J, Stuart DI
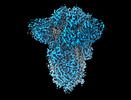
EMDB-18808: 
SD1-3 Fab in complex with SARS-CoV-2 BA.2.12.1 Spike Glycoprotein
Method: single particle / : Duyvesteyn HME, Ren J, Stuart DI

EMDB-41423: 
Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41424: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 1
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41425: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 2
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41777: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:Pomalidomide:SD40
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41778: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 1
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-41779: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 2
Method: single particle / : Roy Burman SS, Hunkeler M, Fischer ES

EMDB-29974: 
Cryo-EM structure of synthetic tetrameric building block sC4
Method: single particle / : Redler RL, Huddy TF, Hsia Y, Baker D, Ekiert D, Bhabha G

EMDB-41364: 
CryoEM Structure of a Computationally Designed T3 Tetrahedral Nanocage
Method: single particle / : Weidle C, Borst AJ

EMDB-42906: 
Computational Designed Nanocage O43_129
Method: single particle / : Weidle C, Kibler RD

EMDB-42944: 
Computational Designed Nanocage O43_129_+4
Method: single particle / : Carr KD, Weidle C, Borst AJ

EMDB-16680: 
BA.4/5-5 FAB IN COMPLEX WITH SARS-COV-2 BA.4 SPIKE GLYCOPROTEIN
Method: single particle / : Duyvesteyn HME, Ren J, Stuart DI, Fry EE

EMDB-35678: 
Apo state of Arabidopsis AZG1 at pH 7.4
Method: single particle / : Xu L, Guo J

EMDB-35679: 
Endogenous substrate adenine bound state of Arabidopsis AZG1 at pH 5.5
Method: single particle / : Xu L, Guo J

EMDB-35680: 
6-BAP bound state of Arabidopsis AZG1
Method: single particle / : Xu L, Guo J

EMDB-35681: 
trans-Zeatin bound state of Arabidopsis AZG1 at pH7.4
Method: single particle / : Xu L, Guo J

EMDB-35682: 
kinetin bound state of Arabidopsis AZG1
Method: single particle / : Xu L, Guo J

EMDB-37658: 
trans-Zeatin bound state of Arabidopsis AZG1 at pH5.5
Method: single particle / : Xu L, Guo J

EMDB-37681: 
Apo state of Arabidopsis AZG1 T440Y
Method: single particle / : Xu L, Guo J
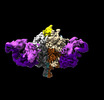
EMDB-41048: 
Lassa GPC Trimer in complex with Fab 8.11G and nanobody D5
Method: single particle / : Gorman J, Kwong PD
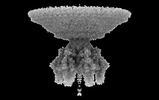
EMDB-41649: 
P22 Mature Virion tail - C6 Localized Reconstruction
Method: single particle / : Iglesias S, Cingolani G, Feng-Hou C
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
